Navigation auf uzh.ch
Navigation auf uzh.ch
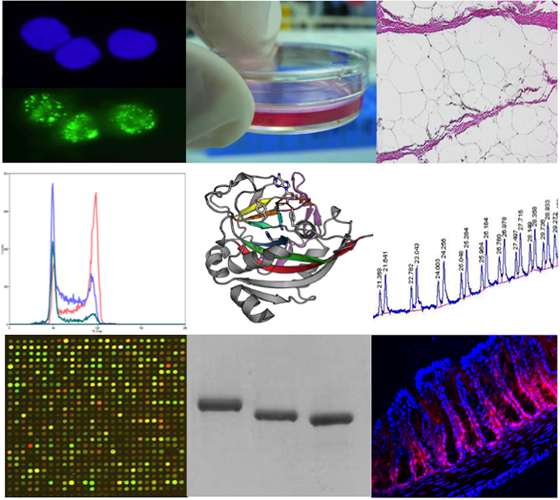
My laboratory is interested to understand the molecular regulatory mechanisms of inflammation. Inflammation is the complex biological response to harmful stimuli, such as pathogens, damaged cells, or irritants. It is a protective attempt by the organism to remove the injurious stimuli as well as to initiate the healing process of the tissue. While inflammation at large is a beneficial event for the organism, excessive activation or inappropriate regulation of immune and inflammation cascades cause tissue and cellular damage, which may lead to cellular dysfunction and cell death.
We investigate inflammatory signaling (e.g. oxidative stress) with special focus on the role of post-translations modifications (PTM) such as ADP-ribosylation in the regulation of inflammation. PTMs of proteins are thought to contribute to the observed complexity of cellular processes in animal and man. PTMs may help to explain the differences between e.g. worm and man, considering that the number and the nucleotide sequences of genes is rather comparable among animal species.
We and others identified protein ADP-ribosylation as a crucial process in the cellular response to detrimental stimuli, be it through genotoxicity, oxidative or metabolic stress, or excessive inflammation. We study the patterns of ADP-ribosylation using cutting-edge systems biology approaches such as ADP-ribosyl-specific high-resolution and quantitative mass spectrometry that we developed in house or in collaboration with various colleagues worldwide. Furthermore, we investigate the players involved, such as writers, readers and eraser of ADP-ribosylation, as well as their target proteins that carry this PTM using state-of-the-art proteomics methods.
Our current work focuses on the following questions: (i) Are ADP-ribosylation patterns cell-type specific or stimulus-specific? I.e. can we delineate specific patterns that are associated with specific cell-physiological conditions? (ii) What are the modifiers (writer, binders, erasers) involved in those patterns? How can we specifically interfere with their activity to modulate the patterns? (iii) What are the regulatory mechanisms in ADP-ribosylation? How is the type and quantity of ADP-ribosylation patterns regulated? (iv) What are the downstream events of ADP-ribosylation? I.e. how does ADP-ribosylation alter the function of specific proteins and what is the effect on cell physiology?